Paper Title: Machine Learning-based integration of multi-omics data for identification of tubular epithelial cell-specific biomarkers in diabetic nephropathy
Authors: Wenning Li, Suriyakala Perumal Chandran
Corresponding Author: Suriyakala Perumal Chandran (suriyakala@lincoln.edu.my)/Malaysia
Abstract
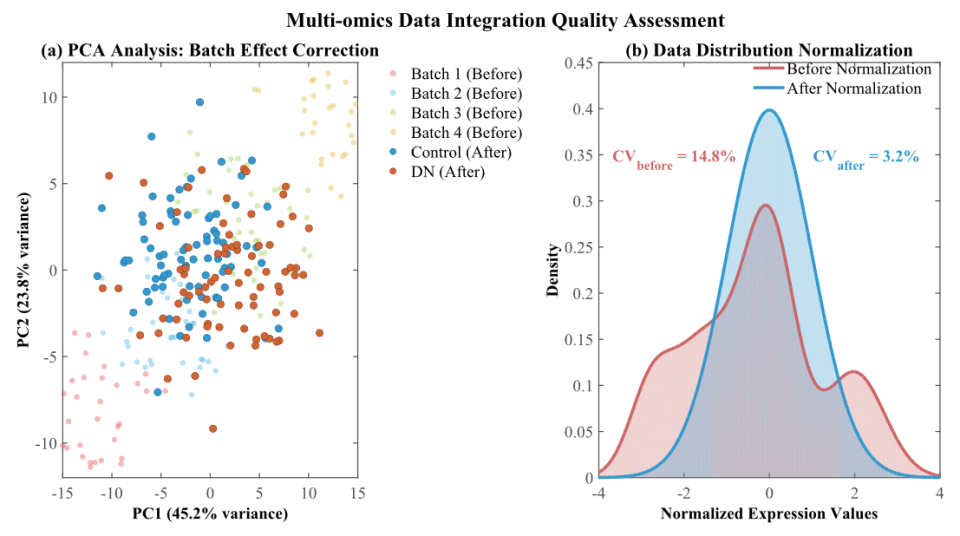
Diabetic nephropathy is a leading cause of end-stage renal disease. Current diagnostic methods, which utilize conventional biomarkers, fail to adequately capture early-stage tubular epithelial cell dysfunction, a condition that likely occurs prior to glomerular damage. This study developed a comprehensive machine learning framework integrating multi-omics data to identify tubular epithelial cell-specific biomarkers for diabetic nephropathy. We systematically collected omics data from established public databases, analyzing 245 transcriptomic samples (18,632 features), 198 proteomic samples (4,521 features), and 167 metabolomic samples (812 features), resulting in an integrated dataset of 156 samples with 23,965 molecular features. Following stringent quality control, batch effect removal, and normalization, we implemented an ensemble learning approach combining Random Forest, Support Vector Machine, and XGBoost algorithms. The ensemble model achieved superior performance with 91.4% accuracy, 89.6% sensitivity, 92.8% specificity, and an AUC of 0.947, representing significant improvement over conventional clinical markers. We identified ten tubular epithelial cell-specific candidate biomarkers, with KIM-1 showing the highest importance score (0.092), followed by NGAL (0.087) and L-FABP (0.084). These markers demonstrated progressive upregulation throughout disease stages with 1.5-fold to 3.2-fold increases in advanced states. Analysis revealed perturbations in inflammatory response pathways, oxidative stress processes, and epithelial-to-mesenchymal transition. Independent cohort validation across three geographically distinct populations confirmed the robustness and generalizability of identified biomarkers. The findings demonstrate the potential of machine learning-based multi-omics integration for enhanced diabetic nephropathy detection and provide novel insights into tubular pathophysiology that could facilitate earlier intervention and personalized treatment strategies.
Keywords
Diabetic nephropathy, Multi-omics integration, Tubular epithelial cells, Machine learning biomarkers, Ensemble algorithms